Full text loading...
A significant challenge for plants is to induce localized defense responses at sites of pathogen attack. Therefore, host subcellular trafficking processes enable accumulation and exchange of defense compounds, which contributes to the plant on-site defenses in response to pathogen perception. This review summarizes our current understanding of the transport processes that facilitate immunity, the significance of which is highlighted by pathogens reprogramming membrane trafficking through host cell translocated effectors. Prominent immune-related cargos of plant trafficking pathways are the pattern recognition receptors (PRRs), which must be present at the plasma membrane to sense microbes in the apoplast. We focus on the dynamic localization of the FLS2 receptor and discuss the pathways that regulate receptor transport within the cell and their link to FLS2-mediated immunity. One emerging theme is that ligand-induced late endocytic trafficking is conserved across different PRR protein families as well as across different plant species.

Article metrics loading...

Full text loading...
Literature Cited


Data & Media loading...
Supplemental Material
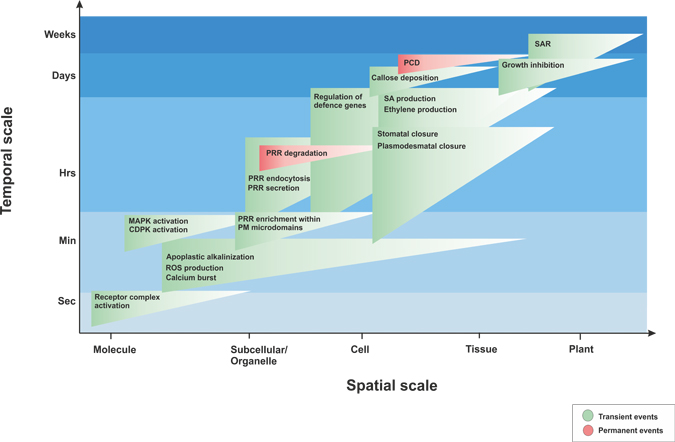
Download all Supplemental Material as a single PDF. Includes: Supplemental Figure 1. Temporal and spatial scales of MAMP-induced events. Known MAMP-induced events are shown according to their time-course of activation upon flg22 elicitation, indicated by blue shading. Triangles indicate the localization of the flg22-induced events. Green colouring indicates transient events, red colouring indicates permanent events.