Full text loading...
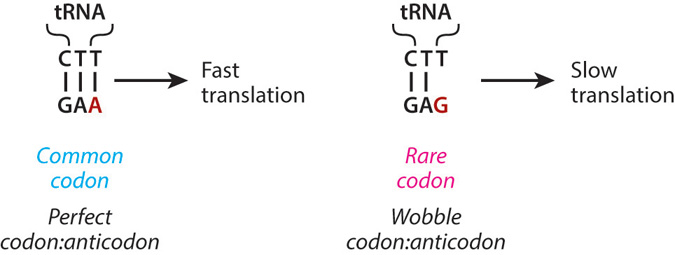
Owing to the degeneracy of the genetic code, a protein sequence can be encoded by many different synonymous mRNA coding sequences. Synonymous codon usage was once thought to be functionally neutral, but evidence now indicates it is shaped by evolutionary selection and affects other aspects of protein biogenesis beyond specifying the amino acid sequence of the protein. Synonymous rare codons, once thought to have only negative impacts on the speed and accuracy of translation, are now known to play an important role in diverse functions, including regulation of cotranslational folding, covalent modifications, secretion, and expression level. Mutations altering synonymous codon usage are linked to human diseases. However, much remains unknown about the molecular mechanisms connecting synonymous codon usage to efficient protein biogenesis and proper cell physiology. Here we review recent literature on the functional effects of codon usage, including bioinformatics approaches aimed at identifying general roles for synonymous codon usage.

Article metrics loading...

Full text loading...
Literature Cited


Data & Media loading...
Supplemental Material
Supplemental Figure 1.